CISC 333 Weka
Tutorial - Part 2
Selecting Attributes
Weka also provides techniques to discard irrelevant attributes and/or reduce the dimensionality of your dataset. After loading a dataset, click on the select attributes tag to open a GUI which will allow you to choose both the evaluation method (such as Principal Components Analysis for example) and the search method (f. ex. greedy or exhaustive search). Depending on the chosen combination, the actual time spend on selecting attributes can vary substantially and can also be very long, even for small datasets such as the Iris data with only five features (including the class attribute) for each of the 150 samples. The picture below shows the results for a sample application. It is also important to note that not all evaluation/search method combinations are valid, watch out for the error message in the Status bar. There's also a problem using Discretize while in the preprocessing mode, which leads to false results. If you need to use this filter, you can work around this by using the FilteredClassifier option in the classify menu.Classification
Clicking on the
classifier tab after loading a dataset into Weka and selecting the
choose tab will bring up a menu with a number of choices for the
classifier that is to be applied to the dataset. Note that you have 4
options on how to test the model you're building: Using the test set, a
training set (you will need to specify the location of the training set
in this case), cross validation and a percentage. The achieved accuracy
of your model will vary, depending on the option you select. One
pitfall to avoid is to select the training set as a test set, as that
will result in an underestimate of the error rate. The resulting
model (with a lot of additional information will be displayed after you
click on start. What exactly is contained in the output can be
determined under options. A sample output for applying the J48 decision
tree algorithm to the Iris dataset is shown in the Figure below.One of the things to watch out for
is that the confusion matrix is displayed, as this gives a lot more
information than just the prediction accuracy. Other useful things are
the options showing up when right clicking the results list on the
bottom right. For example, this is where you can load and save the
models you built, as well as save the results page. Another fact
to keep in mind is that Weka gives hints on how to achieve the same
result from the command line: look at what is displayed next to the
Choose button and how it changes with the options that you select. This
information can also be found towards the top of your results page.
Clustering
The clustering option is very similar to the classification described above, with a few differences regarding the options you select. For instance, there is an easy way to discard undesired attributes.Association Rules
Weka also provides three algorithms to extract association rules from non-numerical data as shown in the picture below.Opening Databases and URL's
Detailed
instructions for loading a windows database into Weka can be found here.
Weka also allows a user to select a URL as the source for a datafile
(click on Open URL from the preprocess pane) or download data from a
database (Open DB from the preprocess pane).The CLI
Sometimes
it's desirable to run Weka from a command line interface, for example
if you want to set up your experiment using a batch file. A
tutorial on how to do this can be found here. This page also
gives instructions on how to call Weka from Java. Anybody
interested in using the CLI should take the time to read through this
tutorial.Examples for using the CLI are :
java weka.filters.unsupervised.attribute.Remove -V -R 1,4 -i trainingFile.arff -o myTrainingFile.arff
This filter removes all but the first and fourth attribute from a dataset stored in a file called trainingFile.arff and stores the result in myTrainingFile.arff
java weka.classifiers.trees.J48 -t myTrainingFile.arff -T myTestFile.arff -U -p 1 > Results.arff
Here, the J48 decision tree is applied to the file myTrainingFile.arff, which was created in the previous step. The test file is specified with the -T option. The results are redirected from the screen to a file called Results.arff and the options -U and -p 1 are options that determine the specific output form.
Experimenter
The experimenter, which can be run from both the command line and a GUI (easier to use), is a tool that allows you to perform more than one experiment at a time, maybe applying different techniques to a datasets, or the same technique repeatedly with different parameters. The Weka homepage provides a link to a tutorial for an earlier version of the Experimenter, which can be downloaded from here.If you choose the experimenter after starting Weka, you get the following screen.
After selecting new, which
initializes a new experiment with default parameters, you can select
where you want to store the results of your experiment by using browse
(there are a number of choices available for the format of your results
file). You can then change the default parameters if desired ( watch
out for the option of selecting classification or regression). For
example, you can add more datasets, delete the ones you already
selected as well as add and delete algorithms applied to your selected
datasets. You can also the type of experiment (cross validation or a
percentage split for the training and test set).
The following picture shows the setup for a n 8 fold cross validation, applying a decision tree and Naive Bayes to the iris and labor dataset that are included in the Weka Package. The results are to be stored in an ARFF file called MyResults.arff in the specified subfolder.
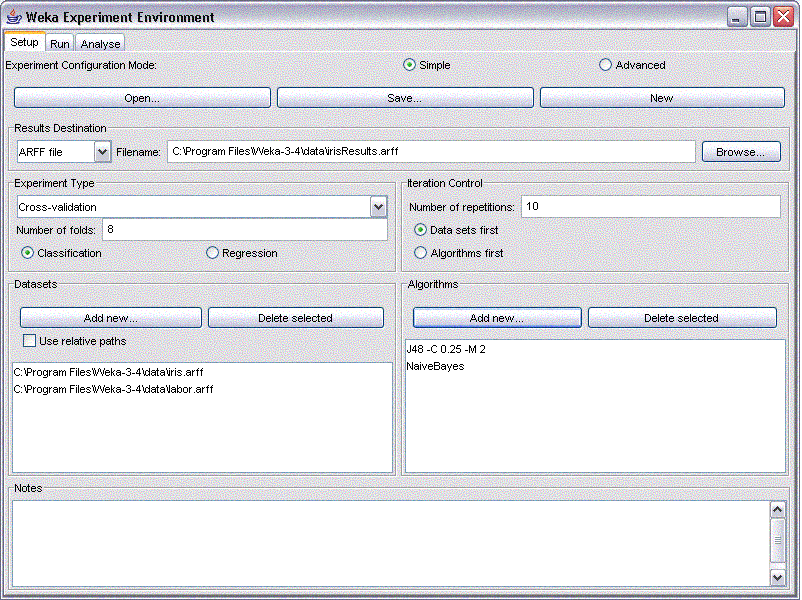
The following picture shows the setup for a n 8 fold cross validation, applying a decision tree and Naive Bayes to the iris and labor dataset that are included in the Weka Package. The results are to be stored in an ARFF file called MyResults.arff in the specified subfolder.
After running your experiment by
selecting Start from the Run tab, your results will be stored in the
specified Results file if the run was successful. You then need to load
this file into Weka from the Analysis pane to see your results. The
picture below shows the Analysis pane after loading the results file
for the experiment set up above.